�ȥåץڡ��� | �ץ��ե����� | �������� | �������� | ���� | ���Υ˥塼�� | ������ | ����¾ | ���λ�Ƴ�������������? | English
�������� †
Drug-Target Interactionͽ¬ †
����ѥ��������ޤΣ����ͥåȥ����ͽ¬���뿷����ˡ��ȯ���Ƥ��ޤ���
BO-DTI †
�٥�����Ŭ�� (GP-MIˡ) �ˤ�ä�DTIͽ¬�γؽ��ˤ�������֤���10�ܹ�®�����ޤ�����
- Tomohiro Ban, Masahito Ohue, Yutaka Akiyama: "Efficient Hyperparameter Optimization by Using Bayesian Optimization for Drug-Target Interaction Prediction", In Proceedings of the 7th IEEE International Conference on Computational Advances in Bio and Medical Sciences (ICCABS 2017), Orlando, FL, USA, October 19-21, 2017. (accepted)
[ Paper | Slide ]
http://www.github.com/akiyamalab/BO-DTI
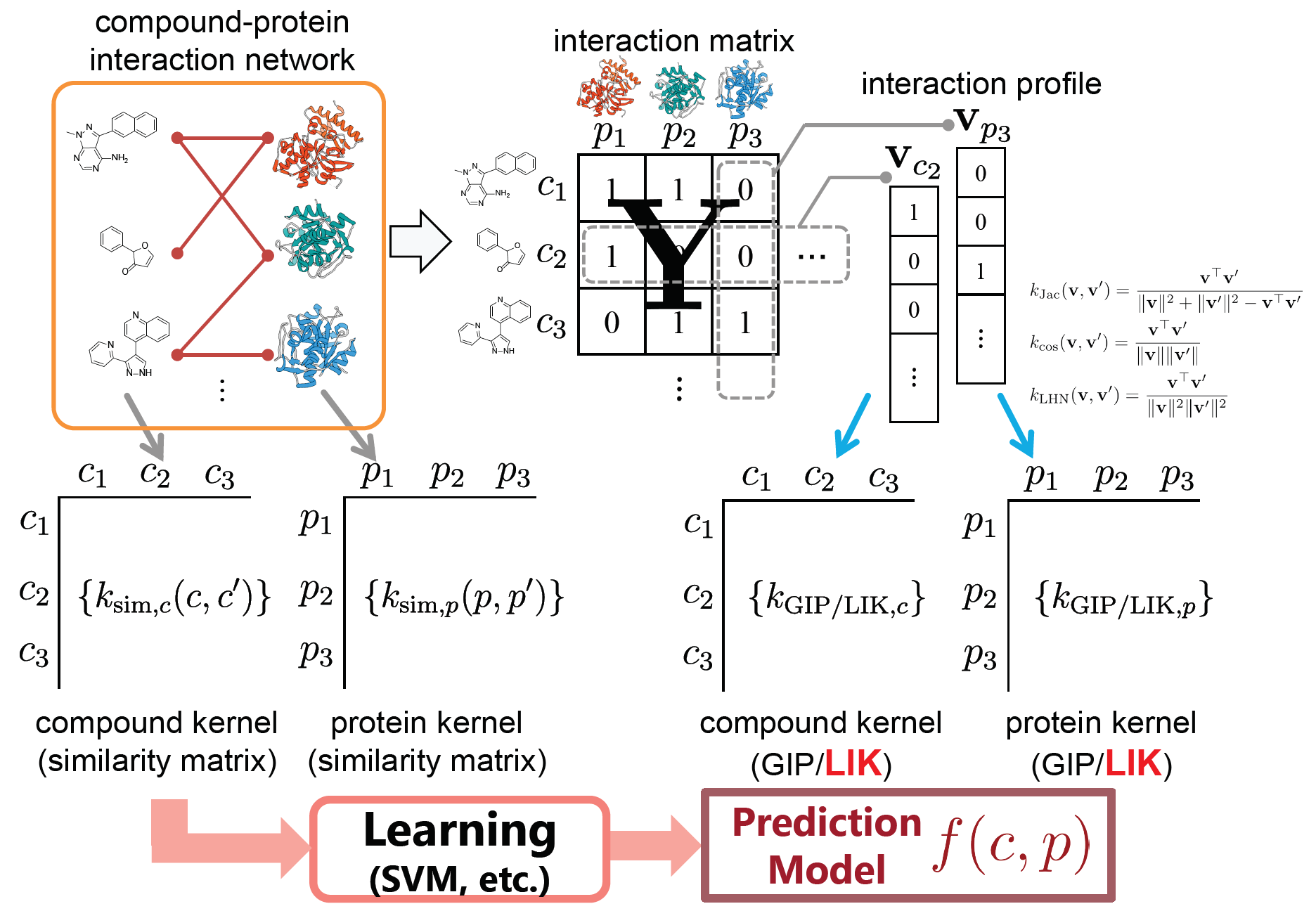
Link Indicator Kernel †
���ɸ���Ѥ����ޥ���ץ륫���ͥ�ˡ����Ƥ��ޤ�����
- Masahito Ohue, Takuro Yamazaki, Tomohiro Ban, Yutaka Akiyama: "Link Mining for Kernel-based Compound-Protein Interaction Predictions Using a Chemogenomics Approach", In Proceedings of the Thirteenth International Conference On Intelligent Computing (ICIC2017) (Lecture Notes in Computer Science), 10362, 549-558, Liverpool,UK August 7-10, 2017.
[ Paper | arXiv | Slide ]
Spresso †
����ѥ����Σ�������¤�˴�Ť�����®�ʥץ졦������˥����ޤ���
http://www.bi.cs.titech.ac.jp/spresso
- Keisuke Yanagisawa, Shunta Komine, Shogo D. Suzuki, Masahito Ohue, Takashi Ishida, Yutaka Akiyama: "Spresso: An ultrafast compound pre-screening method based on compound decomposition", Bioinformatics. (in press)
[ PubMed | Journal Website ]
- Keisuke Yanagisawa, Shunta Komine, Shogo D. Suzuki, Masahito Ohue, Takashi Ishida, Yutaka Akiyama: "ESPRESSO: An ultrafast compound pre-screening method based on compound decomposition", In Proceedings of The 27th International Conference on Genome Informatics (GIW 2016), 2016.
MEGADOCK †

����ѥ�����Ω�ι�¤��������Ѥ��륿��ѥ�������ߺ���ͽ¬�����ƥ�Ǥ���
FFT-base�Υɥå����르�ꥺ���Ƨ�����ʤ���⡤��������ǥ�β��ɤ䥹��å����ʤɤˤ�ä�
��®�ʷ���¸����Ƥ��ޤ���
�ܤ����Ϥ����顧���եȥ�����
NVIDIA Application Catalog�ˤ�Ǻܤ���Ƥ��ޤ���https://www.nvidia.com/en-us/data-center/gpu-accelerated-applications/catalog/
����ѥ����֥ɥå���ͽ¬ †
����ѥ�����Ω�ι�¤���顤�����λ���������Ʊ�Τ��������ʷ����̱��������ˤ�ɽ���Ų٤���ߺ��Ѥʤɤ���Ϥ��ơ�ʣ���Τη����®��ͽ¬������ˡ�椷�Ƥ��ޤ���
- Masahito Ohue†, Yuri Matsuzaki†, Nobuyuki Uchikoga, Takashi Ishida, Yutaka Akiyama: "MEGADOCK: An all-to-all protein-protein interaction prediction system using tertiary structure data", Protein and Peptide Letters, 21(8): 766-778, 2014.
[ PubMed | BenthamScience ]
�������ܥ����륹������ǥ�γ�ȯ †
rPSC�ȸƤФ���ȼ��η�����������������ǥ��١����Ȥ���������ߺ��Ѥ�æ���¼�ͳ���ͥ륮�����ǥ벽��������®�˷���ǽ�ʥ�������ǥ����Ƥ��Ƥ��ޤ���
- Masahito Ohue, Yuri Matsuzaki, Takashi Ishida, Yutaka Akiyama: "Improvement of the Protein-Protein Docking Prediction by Introducing a Simple Hydrophobic Interaction Model: an Application to Interaction Pathway Analysis", In Proceedings of The 7th IAPR International Conference on Pattern Recognition in Bioinformatics (PRIB2012), Lecture Notes in Bioinformatics 7632, 178-187, Springer Heidelberg, 2012.
[ SpringerLink | Slide | Streaming(e-Bio) ]
High Performance Computing (HPC) †
��������ʥ����ѡ�����ԥ塼���ˤ����Ѥ�������ѥ����ɥå���ͽ¬����Ԥ����եȥ�������ȯ���Ƥ��ޤ���
- Masahito Ohue†, Takehiro Shimoda†, Shuji Suzuki, Yuri Matsuzaki, Takashi Ishida, Yutaka Akiyama: "MEGADOCK 4.0: an ultra–high-performance protein–protein docking software for heterogeneous supercomputers", Bioinformatics, 30(22): 3281-3283, 2014.
[ PubMed | Oxford ]
- Yuri Matsuzaki, Nobuyuki Uchikoga, Masahito Ohue, Takehiro Shimoda, Toshiyuki Sato, Takashi Ishida, Yutaka Akiyama: "MEGADOCK 3.0: A high-performance protein-protein interaction prediction software using hybrid parallel computing for petascale supercomputing environments", Source Code for Biology and Medicine, 8(1): 18, 2013.
[ PubMed | BioMedCentral ]
����ѥ�������ߺ��ѥͥåȥ��ͽ¬ †
����ѥ����֤���ߺ��Ѥϡ�����Ǥι���ȿ�����˦�⳰�Ǥξ�����ã�δ��ܤȤʤꡤ��̿��ư�κ�����٤��Ƥ��ޤ����桹�ϥ���ѥ�����Ω�ι�¤�˴�Ť��ɥå������顤��ߺ��Ѥ륿��ѥ����ڥ����®��ͽ¬������ˡ�椷�Ƥ��ޤ���
- Masahito Ohue†, Yuri Matsuzaki†, Nobuyuki Uchikoga, Takashi Ishida, Yutaka Akiyama: "MEGADOCK: An all-to-all protein-protein interaction prediction system using tertiary structure data", Protein and Peptide Letters, 21(8): 766-778, 2014.
[ PubMed | BenthamScience ]
- ������, ����ͳ��, ����͵��, ��ƣ��Ƿ, ������: ��MEGADOCK��Ω�ι�¤���������Ū����ѥ�������ߺ���ͽ¬�Ȥ��Υ����ƥ���ʪ�ؤؤα��ѡ�, ��������ز���ʸ�� ������ǥ벽�ȱ���, 3(3): 91-106, 2010.
[ IPSJ Digital Library ]
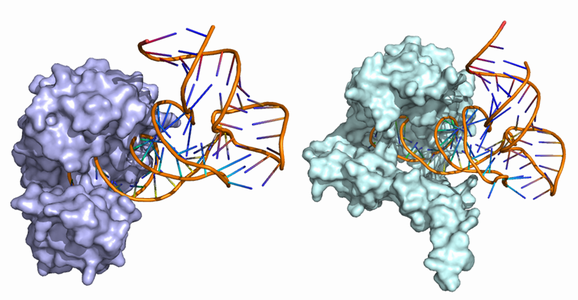
����ѥ���-RNA����ߺ���ͽ¬ †
�ҥȥ��Υ���ˤ�non-coding RNA����1500�Ĥۤɥ����ɤ���Ƥ���Ȥ����Ƥ��ꡤ����ѥ���Ʊ�Τ����Ǥʤ���¿����RNA������ѥ�������ߺ��Ѥ��Ƥ��ޤ����桹���ȼ��Υ���ѥ�������ߺ���ͽ¬�μ�ˡ����Ѥ���������������ѥ���-RNA����ߺ���ͽ¬�μ�ˡ�椷�Ƥ��ޤ���
- Masahito Ohue, Yuri Matsuzaki, Yutaka Akiyama: "Docking-calculation-based Method for Predicting Protein-RNA Interactions",
Genome Informatics, 25(1): 25-39, Aug 3 2011.
[ PubMed | J-stage ]
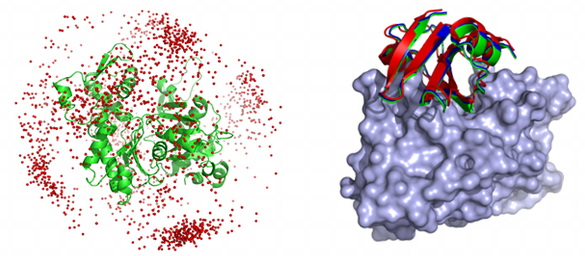
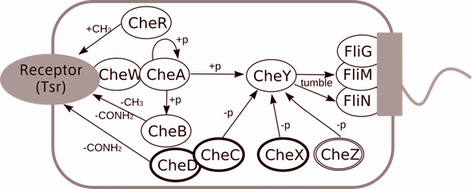
�ٶ��������ѥ����������Ϥؤα��� †
��������λɷ�˱������Ʊ�ư�������������������ȸƤӤޤ���
�㤨�С���IJ�ݤϵ�����֤ˤʤ�ȡ��٤��Ӥΰ����Ҥ�ȯ������
���ܤȤʤ�ʪ���Τ��¿�����ذ�ư���벽���������������ˤ��ޤ���
����������¸����Ƥ���Τϡ��ɷ�˱����ƷϤ��������֤��Ѥ������˦��Υ����ʥ���ã�ϤǤ���
�����ξ��֤��Τ����������֤��Ѥ��Ĥ��б�����Ȥ�����ư�ϡ�
�ؽ��Ȥ�����ǰ�ˤ�Ĥʤ������Ѷ�̣�������ݤǤ���
�ޤ����������Ϥˤ�ä����椵���٤��ӥ⡼�����ϡ�
�����ƥ��ͥ륮����Ψ���ɤ�ʬ�ҥ⡼�����Ȥ��Ƥ����ܤ��졤
����ʬ��ؤα��Ѹ�����Τ��Ƥ��ޤ���
�ܸ���Ǥϡ����ηϤδ��Τ�PPI�������פ��������MEGADOCK��ɾ����Ԥ��ȤȤ�ˡ�
���ηϤ�����˴ؤ��̤�Τ���ߺ��Ѥθ��Ф��ߤޤ�����
- Yuri Matsuzaki†, Masahito Ohue†, Nobuyuki Uchikoga, Yutaka Akiyama: "Protein-protein interaction network prediction by using rigid-body docking tools: application to bacterial chemotaxis", Protein and Peptide Letters, 21(8): 790-798, 2014.
[ PubMed | BenthamScience ]
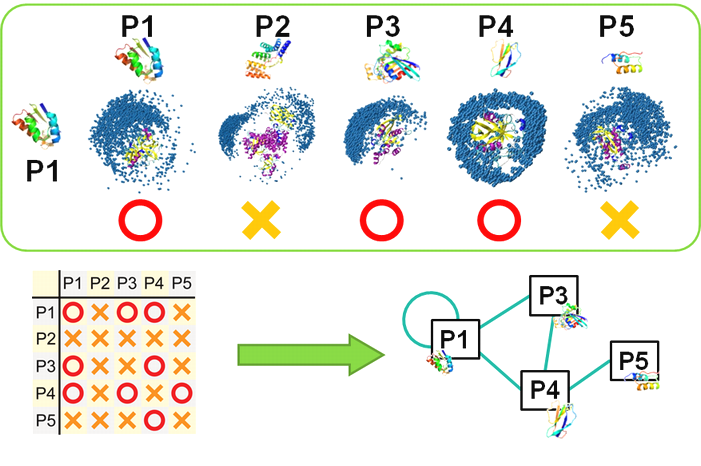
�ҥȥ��ݥȡ������ѥ����������Ϥؤα��� †
���ݥȡ������Ȥϡ����Τξ��֤��ɹ����ݤĤ�����Ѷ�Ū�˰�����������롤
���椵�줿��˦�μ��������Ǥ���
���ݥȡ������θ����˵���������¤Ȥ��ƴ�伫���ȱּ��������ꡤ
���ݥȡ����������ä˵��������ΤȤ��Ƥ�AIDS�����������������졤
�����μ��¤˴ؤ�������������åȤȤ��ơ����ݥȡ������ѥ���������Υ���ѥ����������ܤ���Ƥ��ޤ���
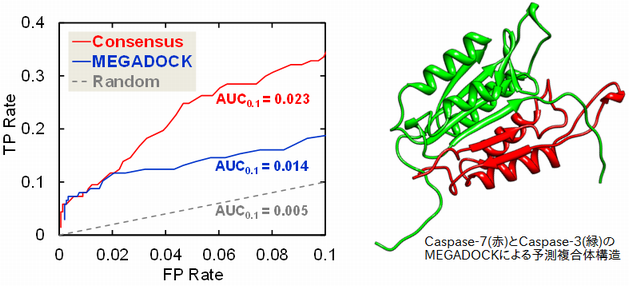
���θ���Ǥϡ�MEGADOCK�ˤ�äƥ��ݥȡ������˴�Ϣ���륿��ѥ�������ߺ��ѥͥåȥ����ͽ¬����ȤȤ�ˡ�����ʣ���ι�¤��ƥ�ץ졼�Ⱦ���Ȥ������Ѥ����ˡ���Ȥ߹�碌�뤳�Ȥ�Ŭ��Ψ�ι⤤ͽ¬���Ԥ��뤳�Ȥ��ޤ�����
- Masahito Ohue, Yuri Matsuzaki, Takehiro Shimoda, Takashi Ishida, Yutaka Akiyama: "Highly Precise Protein-Protein Interaction Prediction Based on Consensus Between Template-Based and de Novo Docking Methods", BMC Proceedings, 7(Suppl 7): S6, 2013.
[ PubMed | BioMedCentral ]
![[PukiWiki] [PukiWiki]](image/pukiwiki.png)
![[PukiWiki] [PukiWiki]](image/pukiwiki.png)